 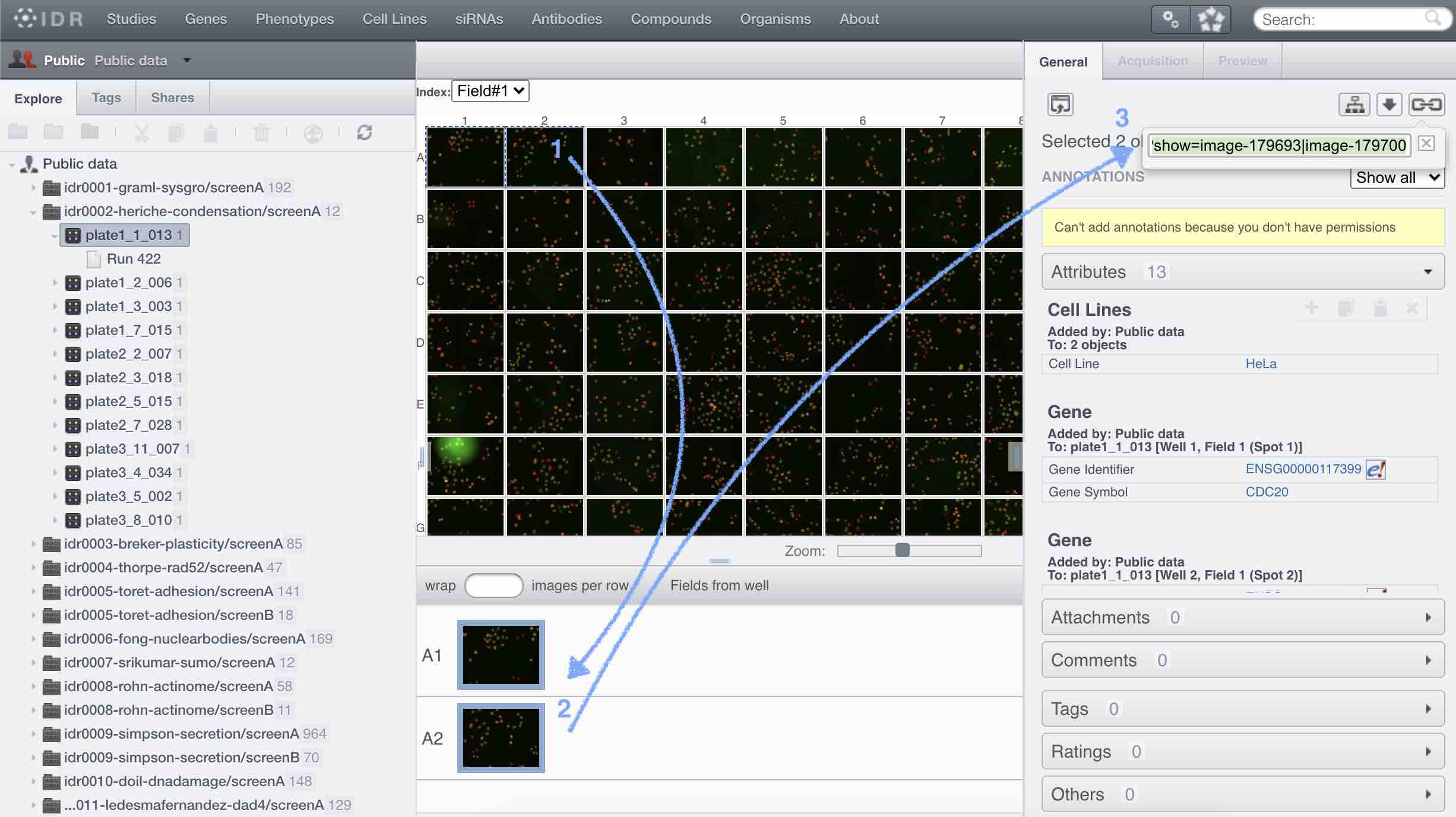
A further validation also demonstrates that previously published results can be reproduced from a publicly available multiplexed image dataset. The efficacy of these operations is demonstrated through several imaging experiments that pair Cytokit results with those from an independent but comparable assay. Image processing operations supported in Cytokit are generally sourced from existing deep learning models or are at least in part adapted from open source packages to run in a single or multi-GPU environment. ResultsĬytokit offers (i) an end-to-end, GPU-accelerated image processing pipeline (ii) efficient input/output (I/O) strategies for operations specific to high dimensional microscopy and (iii) an interactive user interface for cross filtering of spatial, graphical, expression, and morphological cell properties within the 100+ GB image datasets common to multiplexed immunofluorescence. 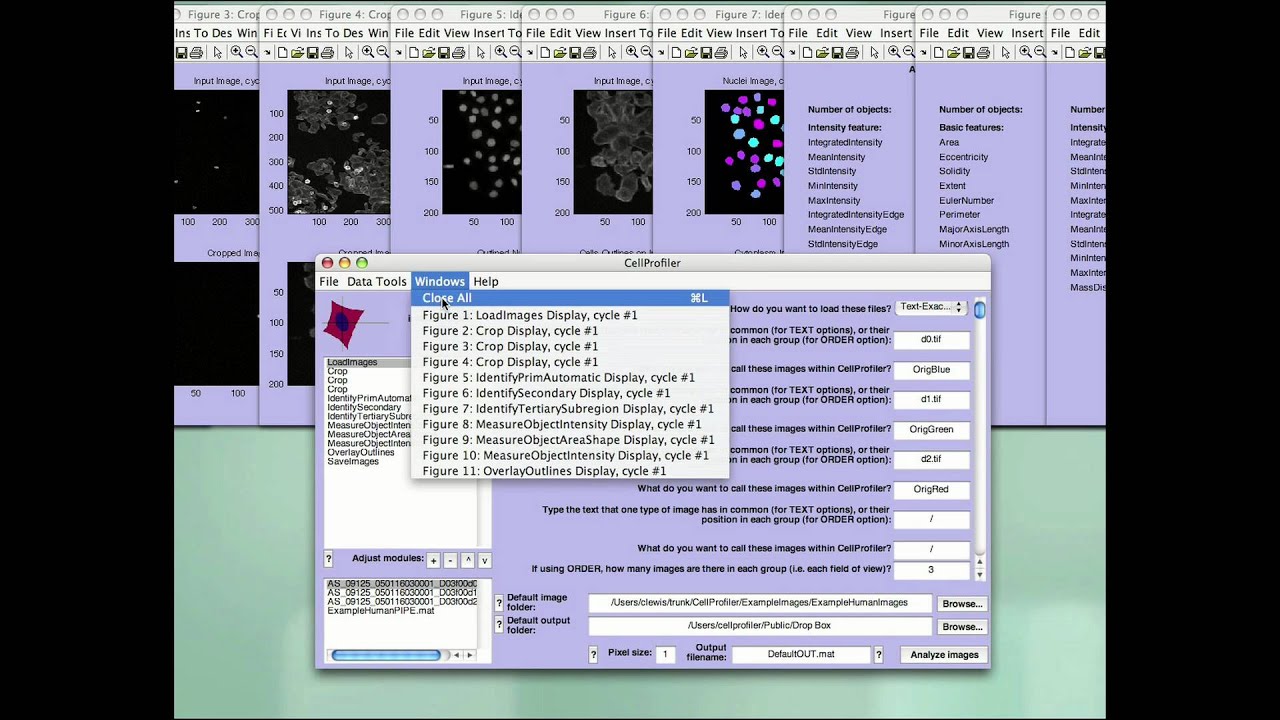 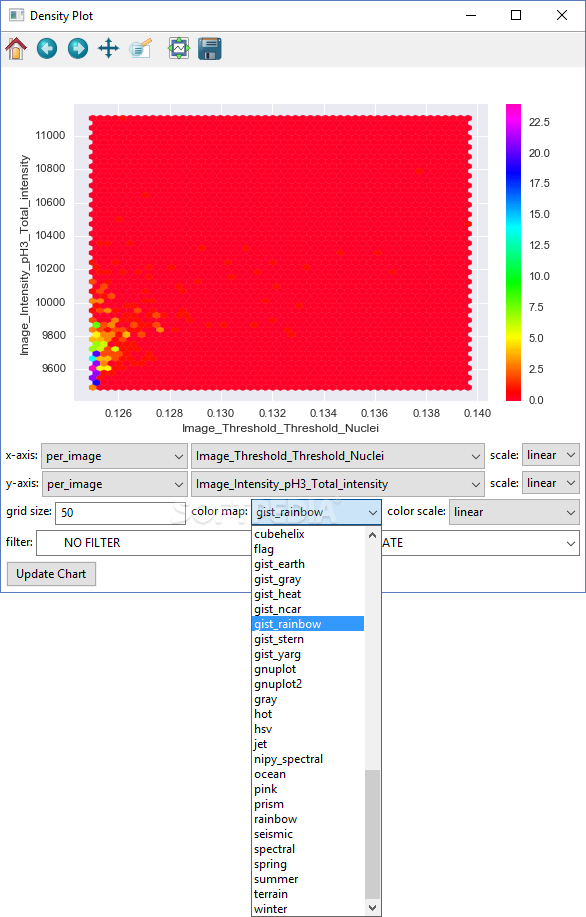
As protocols for conducting such imaging experiments continue to improve, it is the intent of this study to provide and validate software for processing the large quantity of associated data in kind. This spatial information, in addition to morphological properties and extensive intracellular or surface marker profiling, comprise promising avenues for rapid advancements in the understanding of disease progression and diagnosis. Multiplexed in-situ fluorescent imaging offers several advantages over single-cell assays that do not preserve the spatial characteristics of biological samples. 
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
